Multicellular organisms establish and maintain different transcriptional states in disparate cell types through complex and specific regulation of gene expression. This regulation is mediated by the cooperative binding of transcription factors (TFs) to regulatory elements through the recognition of specific DNA sequence motifs. Additionally, the physical access of TFs to DNA can be modulated by epigenetic regulation, such as DNA methylation, nucleosome positioning and histone modifications. Failure to maintain this tight regulation of gene expression results in developmental defects and various diseases including cancer. We use both computational and experimental approaches and integrate genomic data to study TF binding and its regulation in cellular systems.
Transcription factor sensitivity to DNA methylation
Héberlé & Bardet. Essays in Biochemistry (2019)

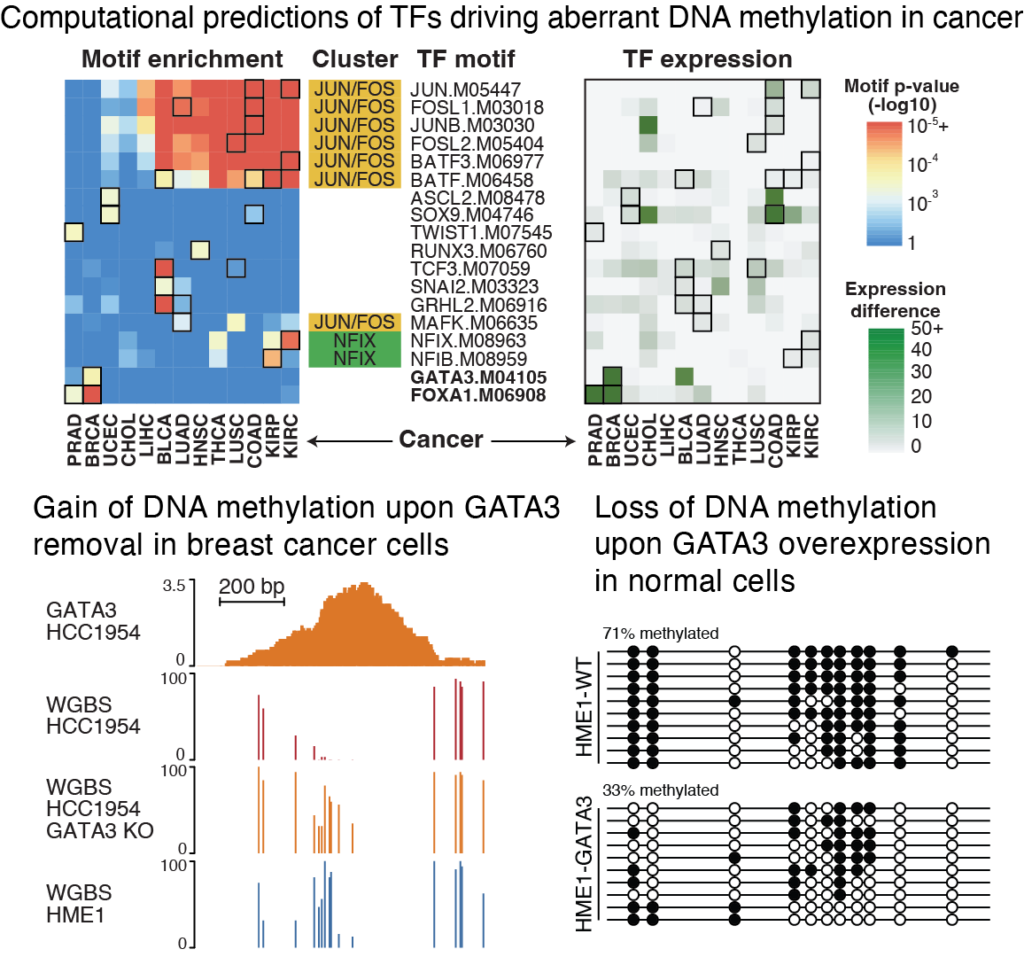
Transcription factor binding in cancer
Detilleux, Spill et al. Epigenetics & Chromatin (2022)
Methods to study transcription factor binding to genomic regions
TFmotifView – Leporcq et al. Nucleic Acid Research (2020)

Methods to study DNA methylation patterns
MethyLasso – Balaramane, Spill et al. BioRxiv (2023)

Funding sources
- Strasbourg University: PhD fellowship to Y. Hadj-Arab
- ANR JCJC (2023-2026, PI)
- IReSP (2023-2027, Partner)
- Ligue Nationale contre le Cancer (2020-2023): PhD fellowship to D. Balaramane
- Plan Cancer: Systems Biology Grant (2018-2023, Coordinator)
- Strasbourg University: IDEX Attractivité Grant (2018-2020, PI)
- CNRS / University of Strasbourg: Basic support
